 |
SqueezeBrains SDK 1.18
|
 |
SqueezeBrains SDK 1.18
|
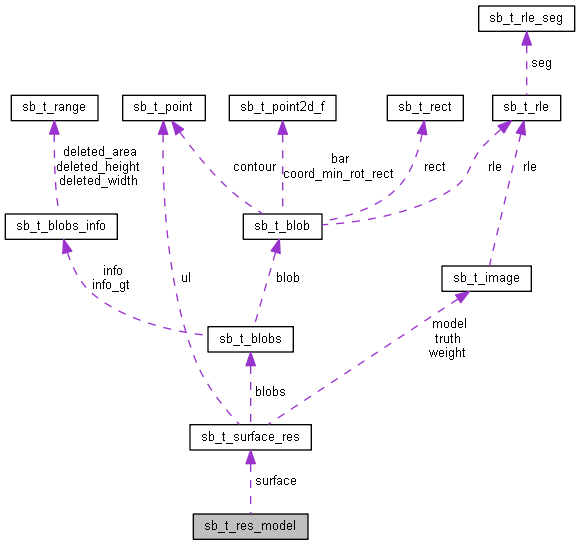
Defines the results of a model. More...
#include <sb.h>
Data Fields | |
| int | tp |
| Count of True Positive. More... | |
| int | tn |
| Count of True Negative. More... | |
| int | fp |
| Count of False Positive. More... | |
| int | fn |
| Count of False Negative. More... | |
| int | op |
| Count of Optional Positive. More... | |
| int | on |
| Count of Optional Negative. More... | |
| int | out_of_roi |
| Count of Out Of ROI. More... | |
| int | disabled |
| Count of samples with model disabled. More... | |
| sb_t_surface_res * | surface |
| Surface model analysis result. More... | |
Defines the results of a model.
The fields tp, tn, fp, fn, op, on, out_of_roi, disabled are evaluated only if the samples, in case of Retina and Deep Cortex project, or roi_defects, in case of Surface and Deep Surface project, are passed to the function sb_project_detection.
There are special cases where tn = 1 is set.
For Surface and Deep Surface projects the results is tn=1 in case of:
For Retina projects the result is tn=1 in case of:
With Retina it is possible to have occurrences classified as True Negative if parameter sb_t_par_model::num_occurrences is set to a value lower than 0.
For Deep Cortex projects the result is tn=1 in case of:
The presence of op or on or or does not influence the tn classification.
| int sb_t_res_model::disabled |
Count of samples with model disabled.
| int sb_t_res_model::fn |
Count of False Negative.
| int sb_t_res_model::fp |
Count of False Positive.
| int sb_t_res_model::on |
Count of Optional Negative.
| int sb_t_res_model::op |
Count of Optional Positive.
| int sb_t_res_model::out_of_roi |
Count of Out Of ROI.
| sb_t_surface_res* sb_t_res_model::surface |
Surface model analysis result.
In case of sb_t_res::models , the field is allocated only if the function sb_project_get_res is called with the parameter details is set to 1.
| int sb_t_res_model::tn |
Count of True Negative.
| int sb_t_res_model::tp |
Count of True Positive.